Agglomerate data using taxonomic information or other grouping
Source:R/AllGenerics.R, R/agglomerate.R, R/splitByRanks.R
agglomerate-methods.RdAgglomeration functions can be used to sum-up data based on specific criteria such as taxonomic ranks, variables or prevalence.
agglomerateByRank can be used to sum up data based on associations
with certain taxonomic ranks, as defined in rowData. Only available
taxonomyRanks can be used.
agglomerateByVariable and agglomerateByModule merge data on
rows or columns of a SummarizedExperiment as defined by a
factor alongside the chosen dimension. This function allows
agglomeration of data based on other variables than taxonomy ranks.
Metadata from the rowData or colData are
retained as defined by archetype.
assay are
agglomerated, i.e. summed up. If the assay contains values other than counts
or absolute values, this can lead to meaningless values being produced.
agglomerateByRanks takes a SummarizedExperiment, splits it
along the
taxonomic ranks, aggregates the data per rank, converts the input to a
SingleCellExperiment objects and stores the aggregated data as
alternative experiments. unsplitByRanks takes these alternative
experiments and flattens them again into a single
SummarizedExperiment.
agglomerateByRank(x, ...)
agglomerateByVariable(x, ...)
agglomerateByModule(x, ...)
agglomerateByRanks(x, ...)
unsplitByRanks(x, ...)
# S4 method for class 'TreeSummarizedExperiment'
agglomerateByRank(
x,
rank = taxonomyRanks(x)[1],
update.tree = agglomerateTree,
agglomerate.tree = agglomerateTree,
agglomerateTree = TRUE,
...
)
# S4 method for class 'SingleCellExperiment'
agglomerateByRank(
x,
rank = taxonomyRanks(x)[1],
altexp = NULL,
altexp.rm = strip_altexp,
strip_altexp = TRUE,
...
)
# S4 method for class 'SummarizedExperiment'
agglomerateByRank(
x,
rank = taxonomyRanks(x)[1],
empty.rm = TRUE,
empty.fields = c(NA, "", " ", "\t", "-", "_"),
...
)
# S4 method for class 'TreeSummarizedExperiment'
agglomerateByVariable(
x,
by,
group = f,
f,
update.tree = mergeTree,
mergeTree = TRUE,
...
)
# S4 method for class 'SummarizedExperiment'
agglomerateByVariable(x, by, group = f, f, ...)
# S4 method for class 'SummarizedExperiment'
agglomerateByModule(x, by, group, na.rm = FALSE)
# S4 method for class 'SummarizedExperiment'
agglomerateByRanks(
x,
ranks = taxonomyRanks(x),
na.rm = TRUE,
as.list = FALSE,
...
)
# S4 method for class 'SingleCellExperiment'
agglomerateByRanks(
x,
ranks = taxonomyRanks(x),
na.rm = TRUE,
as.list = FALSE,
...
)
# S4 method for class 'TreeSummarizedExperiment'
agglomerateByRanks(
x,
ranks = taxonomyRanks(x),
na.rm = TRUE,
as.list = FALSE,
...
)
splitByRanks(x, ...)
# S4 method for class 'SingleCellExperiment'
unsplitByRanks(
x,
ranks = taxonomyRanks(x),
keep.dimred = keep_reducedDims,
keep_reducedDims = FALSE,
...
)
# S4 method for class 'TreeSummarizedExperiment'
unsplitByRanks(
x,
ranks = taxonomyRanks(x),
keep.dimred = keep_reducedDims,
keep_reducedDims = FALSE,
...
)Arguments
- x
- ...
arguments passed to
agglomerateByRankfunction forSummarizedExperimentobjects and other functions. SeeagglomerateByRankfor more details.- rank
Character scalar. Defines a taxonomic rank. Must be a value oftaxonomyRanks()function.- update.tree
Logical scalar. ShouldrowTree()also be merged? (Default:TRUE)- agglomerate.tree
Deprecated. Use
update.treeinstead.- agglomerateTree
Deprecated. Use
update.treeinstead.- altexp
Character scalarorinteger scalar. Specifies an alternative experiment containing the input data.- altexp.rm
Logical scalar. Should alternative experiments be removed prior to agglomeration? This prevents too many nested alternative experiments by default. (Default:TRUE)- strip_altexp
Deprecated. Use
altexp.rminstead.- empty.rm
Logical scalar. Defines whether rows includingempty.fieldsin specifiedrankwill be excluded. (Default:TRUE)- empty.fields
Character vector. Defines which values should be regarded as empty. (Default:c(NA, "", " ", "\t")). They will be removed ifna.rm = TRUEbefore agglomeration.- by
Character scalar. Determines if data is merged row-wise / for features ('rows') or column-wise / for samples ('cols'). Must be'rows'or'cols'.- group
Character scalar,character vectororfactor vector. A column name fromrowData(x)orcolData(x)or alternatively a vector specifying how the merging is performed. If vector, the value must be the same length asnrow(x)/ncol(x). Rows or columns corresponding to the same level will be merged. Iflength(levels(group)) == nrow(x)/ncol(x),xwill be returned unchanged. ForagglomerateByModule,groupshould specify one or several names of numeric or logical variables from therowData(x)/colData(x)by which to agglomerate rows or columns.- f
Deprecated. Use
groupinstead.- mergeTree
Deprecated. Use
update.treeinstead.- na.rm
Logical scalar. Should NA values be omitted? (Default:TRUE)- ranks
Character vector. Defines taxonomic ranks. Must all be values oftaxonomyRanks()function.- as.list
Logical scalar. Should the list ofSummarizedExperimentobjects be returned by the functionagglomerateByRanksas a SimpleList or stored in altExps? (Default:FALSE)- keep.dimred
Logical scalar. Should thereducedDims(x)be transferred to the result? Please note, that this breaks the link between the data used to calculate the reduced dims. (Default:FALSE)- keep_reducedDims
Deprecated. Use
keep.dimredinstead.
Value
agglomerateByRank returns a taxonomically-agglomerated,
optionally-pruned object of the same class as x.
agglomerateByVariable and agglomerateByModule return an object
of the same class as x with the specified entries merged into one
entry in all relevant components.
For agglomerateByRanks:
If as.list = TRUE : SummarizedExperiment objects in a
SimpleList
If as.list = FALSE : The SummarizedExperiment passed as a
parameter and now containing the SummarizedExperiment objects in its
altExps
For unsplitByRanks: x, with rowData and assay
data replaced by the unsplit data. colData of x is kept as well
and any existing rowTree is dropped as well, since existing
rowLinks are not valid anymore.
Details
Agglomeration sums up the values of assays at the specified taxonomic level. With certain assays, e.g. those that include binary or negative values, this summing can produce meaningless values. In those cases, consider performing agglomeration first, and then applying the transformation afterwards.
agglomerateByVariable works similarly to
aggregateAcrossGenes.
However, additional support for TreeSummarizedExperiment was added and
science field agnostic names were used. In addition the archetype
argument lets the user select how to preserve row or column data. For merge
data of assays the function from scuttle are used.
agglomerateByModule allows to agglomerate features or samples based
on one or multiple variables of numeric or logical type. It is particularly
useful for agglomerating by taxonomic or functional modules, each defined by
a logical or binary variable in the rowData, as features can belong to
several modules.
agglomerateByRanks will use by default all available taxonomic ranks,
but
this can be controlled by setting ranks manually. NA values
are removed by default, since they would not make sense, if the result
should be used for unsplitByRanks at some point. The input data
remains unchanged in the returned SingleCellExperiment objects.
unsplitByRanks will remove any NA value on each taxonomic rank
so that no ambiguous data is created. In additional, a column
taxonomicLevel is created or overwritten in the rowData to
specify from which alternative experiment this originates from. This can also
be used for splitAltExps to
split the result along the same factor again. The input data from the base
objects is not returned, only the data from the altExp(). Be aware
that
changes to rowData of the base object are not returned, whereas only
the colData of the base object is kept.
See also
splitOn
unsplitOn
agglomerateByVariable,
agglomerateByRank,
altExps,
splitAltExps
Examples
### Agglomerate data based on taxonomic information
data(GlobalPatterns)
# print the available taxonomic ranks
colnames(rowData(GlobalPatterns))
#> [1] "Kingdom" "Phylum" "Class" "Order" "Family" "Genus" "Species"
taxonomyRanks(GlobalPatterns)
#> [1] "Kingdom" "Phylum" "Class" "Order" "Family" "Genus" "Species"
# agglomerate at the Family taxonomic rank
x1 <- agglomerateByRank(GlobalPatterns, rank="Family")
#> Duplicated labels were made unique.
## How many taxa before/after agglomeration?
nrow(GlobalPatterns)
#> [1] 19216
nrow(x1)
#> [1] 341
# Do not agglomerate the tree
x2 <- agglomerateByRank(
GlobalPatterns, rank="Family", update.tree = FALSE)
#> Duplicated labels were made unique.
nrow(x2) # same number of rows, but
#> [1] 341
rowTree(x1) # ... different
#>
#> Phylogenetic tree with 341 tips and 340 internal nodes.
#>
#> Tip labels:
#> Sulfolobaceae, SAGMA-X, Cenarchaeaceae, Nitrososphaeraceae, Halobacteriaceae, Methanosaetaceae, ...
#> Node labels:
#> , 0.858.4, 0.764.3, 0.985.6, 1.000.112, 0.978.18, ...
#>
#> Rooted; includes branch length(s).
rowTree(x2) # ... tree
#>
#> Phylogenetic tree with 19216 tips and 19215 internal nodes.
#>
#> Tip labels:
#> 549322, 522457, 951, 244423, 586076, 246140, ...
#> Node labels:
#> , 0.858.4, 1.000.154, 0.764.3, 0.995.2, 1.000.2, ...
#>
#> Rooted; includes branch length(s).
# If assay contains binary or negative values, summing might lead to
# meaningless values, and you will get a warning. In these cases, you might
# want to do agglomeration again at chosen taxonomic level.
tse <- transformAssay(GlobalPatterns, method = "pa")
tse <- agglomerateByRank(tse, rank = "Genus")
#> Warning: 'pa' includes binary values.
#> Agglomeration of it might lead to meaningless values.
#> Check the assay, and consider doing transformation againmanually with agglomerated data.
#> Duplicated labels were made unique.
tse <- transformAssay(tse, method = "pa")
# Removing empty labels by setting empty.rm = TRUE
sum(is.na(rowData(GlobalPatterns)$Family))
#> [1] 5603
x3 <- agglomerateByRank(GlobalPatterns, rank="Family", empty.rm = TRUE)
#> Duplicated labels were made unique.
nrow(x3) # different from x2
#> [1] 341
# Because all the rownames are from the same rank, rownames do not include
# prefixes, in this case "Family:".
print(rownames(x3[1:3,]))
#> [1] "125ds10" "211ds20" "5B-12"
# To add them, use getTaxonomyLabels function.
rownames(x3) <- getTaxonomyLabels(x3, with.rank = TRUE)
#> Duplicated labels were made unique.
print(rownames(x3[1:3,]))
#> [1] "Family:125ds10" "Family:211ds20" "Family:5B-12"
# use 'empty.ranks.rm' to remove columns that include only NAs
x4 <- agglomerateByRank(
GlobalPatterns, rank="Phylum", empty.ranks.rm = TRUE)
head(rowData(x4))
#> DataFrame with 6 rows and 2 columns
#> Kingdom Phylum
#> <character> <character>
#> ABY1_OD1 Bacteria ABY1_OD1
#> AC1 Bacteria AC1
#> AD3 Bacteria AD3
#> Acidobacteria Bacteria Acidobacteria
#> Actinobacteria Bacteria Actinobacteria
#> Armatimonadetes Bacteria Armatimonadetes
# If the assay contains NAs, you might want to specify na.rm=TRUE,
# since summing-up NAs lead to NA
x5 <- GlobalPatterns
# Replace first value with NA
assay(x5)[1,1] <- NA
x6 <- agglomerateByRank(x5, "Kingdom")
head( assay(x6) )
#> CL3 CC1 SV1 M31Fcsw M11Fcsw M31Plmr M11Plmr F21Plmr M31Tong
#> Archaea NA 1248 28811 33 57 42 112 140 303
#> Bacteria 862815 1134209 668698 1543418 2076419 718901 433782 186157 2000099
#> M11Tong LMEpi24M SLEpi20M AQC1cm AQC4cm AQC7cm NP2 NP3
#> Archaea 30 131 145 4459 24692 28051 1826 43197
#> Bacteria 100157 2117461 1217167 1163289 2332489 1671242 521808 1435768
#> NP5 TRRsed1 TRRsed2 TRRsed3 TS28 TS29 Even1 Even2 Even3
#> Archaea 33996 843 8418 14250 1598 1690 150 23 91
#> Bacteria 1618758 57845 484708 265454 935868 1209381 1215987 971050 1078150
# Use na.rm=TRUE
x6 <- agglomerateByRank(x5, "Kingdom", na.rm = TRUE)
head( assay(x6) )
#> CL3 CC1 SV1 M31Fcsw M11Fcsw M31Plmr M11Plmr F21Plmr M31Tong
#> Archaea 1262 1248 28811 33 57 42 112 140 303
#> Bacteria 862815 1134209 668698 1543418 2076419 718901 433782 186157 2000099
#> M11Tong LMEpi24M SLEpi20M AQC1cm AQC4cm AQC7cm NP2 NP3
#> Archaea 30 131 145 4459 24692 28051 1826 43197
#> Bacteria 100157 2117461 1217167 1163289 2332489 1671242 521808 1435768
#> NP5 TRRsed1 TRRsed2 TRRsed3 TS28 TS29 Even1 Even2 Even3
#> Archaea 33996 843 8418 14250 1598 1690 150 23 91
#> Bacteria 1618758 57845 484708 265454 935868 1209381 1215987 971050 1078150
## Look at enterotype dataset...
data(enterotype)
## Print the available taxonomic ranks. Shows only 1 available rank,
## not useful for agglomerateByRank
taxonomyRanks(enterotype)
#> [1] "Genus"
### Merge TreeSummarizedExperiments on rows and columns
data(esophagus)
esophagus
#> class: TreeSummarizedExperiment
#> dim: 58 3
#> metadata(0):
#> assays(1): counts
#> rownames(58): 59_8_22 59_5_13 ... 65_9_9 59_2_6
#> rowData names(0):
#> colnames(3): B C D
#> colData names(0):
#> reducedDimNames(0):
#> mainExpName: NULL
#> altExpNames(0):
#> rowLinks: a LinkDataFrame (58 rows)
#> rowTree: 1 phylo tree(s) (58 leaves)
#> colLinks: NULL
#> colTree: NULL
plot(rowTree(esophagus))
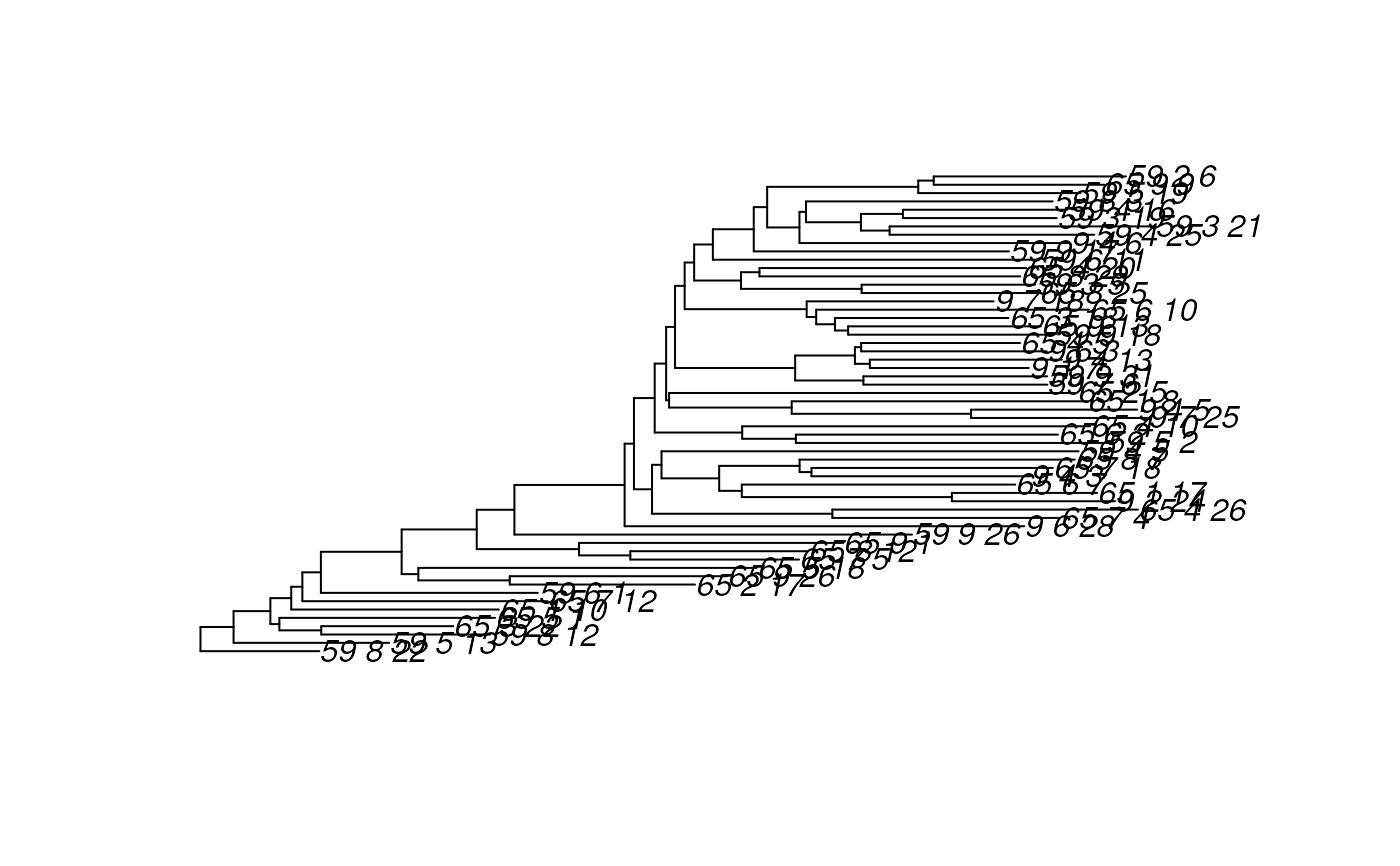 # Get a factor for merging
f <- factor(regmatches(rownames(esophagus),
regexpr("^[0-9]*_[0-9]*",rownames(esophagus))))
merged <- agglomerateByVariable(
esophagus, by = "rows", f, update.tree = TRUE)
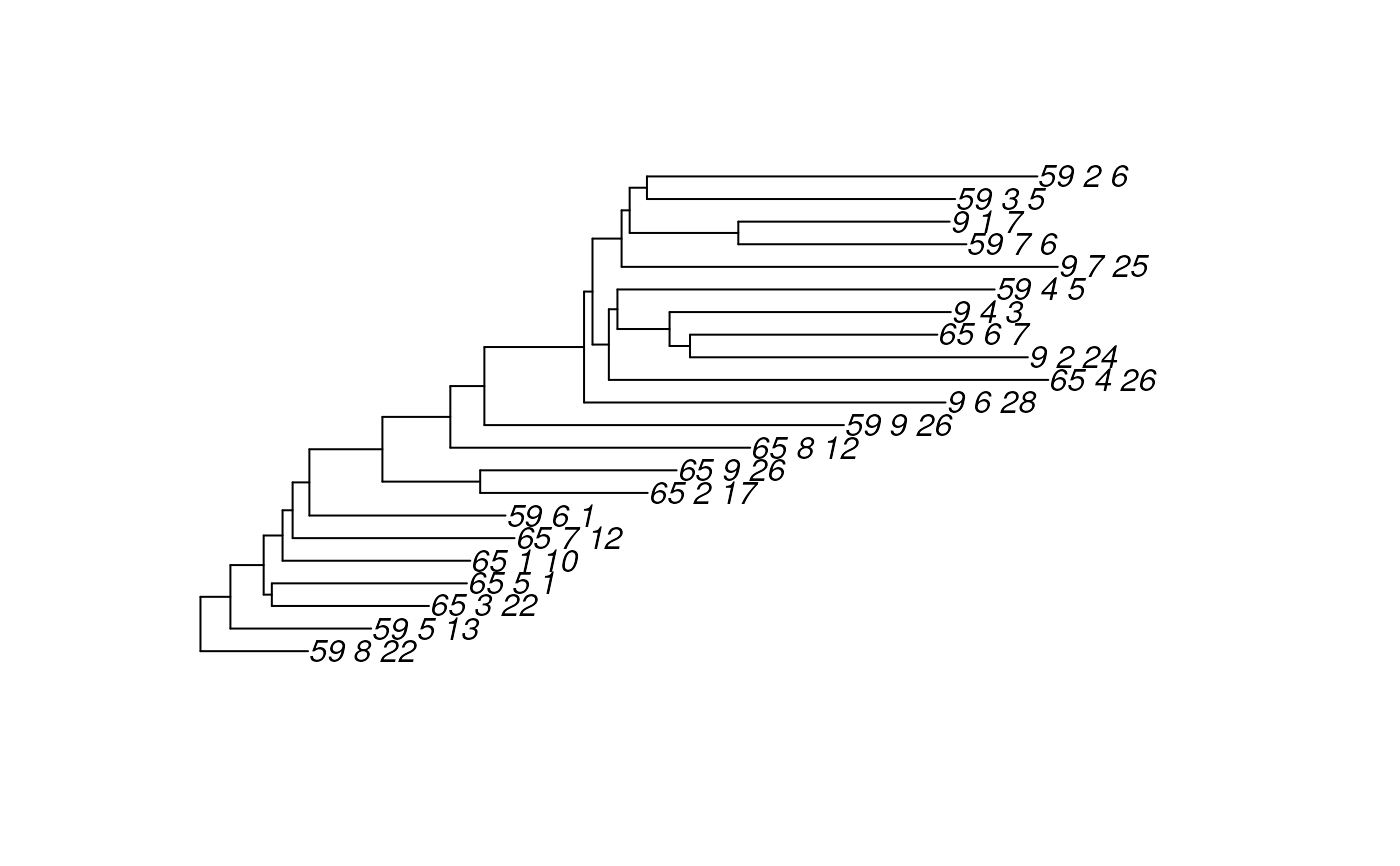
plot(rowTree(merged))
# Get a factor for merging
f <- factor(regmatches(rownames(esophagus),
regexpr("^[0-9]*_[0-9]*",rownames(esophagus))))
merged <- agglomerateByVariable(
esophagus, by = "rows", f, update.tree = TRUE)
plot(rowTree(merged))
 #
data(GlobalPatterns)
GlobalPatterns
#> class: TreeSummarizedExperiment
#> dim: 19216 26
#> metadata(0):
#> assays(1): counts
#> rownames(19216): 549322 522457 ... 200359 271582
#> rowData names(7): Kingdom Phylum ... Genus Species
#> colnames(26): CL3 CC1 ... Even2 Even3
#> colData names(7): X.SampleID Primer ... SampleType Description
#> reducedDimNames(0):
#> mainExpName: NULL
#> altExpNames(0):
#> rowLinks: a LinkDataFrame (19216 rows)
#> rowTree: 1 phylo tree(s) (19216 leaves)
#> colLinks: NULL
#> colTree: NULL
merged <- agglomerateByVariable(
GlobalPatterns, by = "cols", colData(GlobalPatterns)$SampleType)
merged
#> class: TreeSummarizedExperiment
#> dim: 19216 9
#> metadata(0):
#> assays(1): counts
#> rownames(19216): 549322 522457 ... 200359 271582
#> rowData names(7): Kingdom Phylum ... Genus Species
#> colnames(9): Feces Freshwater ... Soil Tongue
#> colData names(7): X.SampleID Primer ... SampleType Description
#> reducedDimNames(0):
#> mainExpName: NULL
#> altExpNames(0):
#> rowLinks: a LinkDataFrame (19216 rows)
#> rowTree: 1 phylo tree(s) (19216 leaves)
#> colLinks: NULL
#> colTree: NULL
## Agglomerate by multiple modules
# Generate 30 random modules
N_module <- 30L
modules <- sample(
c(TRUE, FALSE),
size = nrow(tse) * N_module,
prob = c(0.2, 0.8),
replace = TRUE
)
# Convert modules to matrix
modules <- matrix(modules, nrow = nrow(tse))
# Add module names as colnames
colnames(modules) <- paste0("module_", seq_len(ncol(modules)))
# Add modules to rowData
rowData(tse) <- cbind(rowData(tse), modules)
# Extract module columns
module_columns <- grep("module_", colnames(rowData(tse)), value = TRUE)
# Agglomerate based on modules
tse_module <- agglomerateByModule(tse, by = 1, group = module_columns)
#> Warning: 'pa' includes binary values.
#> Agglomeration of it might lead to meaningless values.
#> Check the assay, and consider doing transformation againmanually with agglomerated data.
# Optionally, store results into altExp slot
altExp(tse, "modules") <- tse_module
data(GlobalPatterns)
# print the available taxonomic ranks
taxonomyRanks(GlobalPatterns)
#> [1] "Kingdom" "Phylum" "Class" "Order" "Family" "Genus" "Species"
# agglomerateByRanks
#
tse <- agglomerateByRanks(GlobalPatterns)
#> Duplicated labels were made unique.
#> Duplicated labels were made unique.
altExps(tse)
#> List of length 7
#> names(7): Kingdom Phylum Class Order Family Genus Species
altExp(tse,"Kingdom")
#> class: TreeSummarizedExperiment
#> dim: 2 26
#> metadata(1): agglomerated_by_rank
#> assays(1): counts
#> rownames(2): Archaea Bacteria
#> rowData names(7): Kingdom Phylum ... Genus Species
#> colnames(26): CL3 CC1 ... Even2 Even3
#> colData names(7): X.SampleID Primer ... SampleType Description
#> reducedDimNames(0):
#> mainExpName: NULL
#> altExpNames(0):
#> rowLinks: a LinkDataFrame (2 rows)
#> rowTree: 1 phylo tree(s) (2 leaves)
#> colLinks: NULL
#> colTree: NULL
altExp(tse,"Species")
#> class: TreeSummarizedExperiment
#> dim: 944 26
#> metadata(1): agglomerated_by_rank
#> assays(1): counts
#> rownames(944): Abiotrophiadefectiva Achromatiumoxaliferum ...
#> proteobacteriumsymbiontofOsedaxsp.MB4 symbiontofNoeetapupillata
#> rowData names(7): Kingdom Phylum ... Genus Species
#> colnames(26): CL3 CC1 ... Even2 Even3
#> colData names(7): X.SampleID Primer ... SampleType Description
#> reducedDimNames(0):
#> mainExpName: NULL
#> altExpNames(0):
#> rowLinks: a LinkDataFrame (944 rows)
#> rowTree: 1 phylo tree(s) (944 leaves)
#> colLinks: NULL
#> colTree: NULL
# unsplitByRanks
tse <- unsplitByRanks(tse)
tse
#> class: TreeSummarizedExperiment
#> dim: 2692 26
#> metadata(0):
#> assays(1): counts
#> rownames(2692): Kingdom:Archaea Kingdom:Bacteria ...
#> Species:proteobacteriumsymbiontofOsedaxsp.MB4
#> Species:symbiontofNoeetapupillata
#> rowData names(8): Kingdom Phylum ... Species taxonomicLevel
#> colnames(26): CL3 CC1 ... Even2 Even3
#> colData names(7): X.SampleID Primer ... SampleType Description
#> reducedDimNames(0):
#> mainExpName: NULL
#> altExpNames(0):
#> rowLinks: NULL
#> rowTree: NULL
#> colLinks: NULL
#> colTree: NULL
#
data(GlobalPatterns)
GlobalPatterns
#> class: TreeSummarizedExperiment
#> dim: 19216 26
#> metadata(0):
#> assays(1): counts
#> rownames(19216): 549322 522457 ... 200359 271582
#> rowData names(7): Kingdom Phylum ... Genus Species
#> colnames(26): CL3 CC1 ... Even2 Even3
#> colData names(7): X.SampleID Primer ... SampleType Description
#> reducedDimNames(0):
#> mainExpName: NULL
#> altExpNames(0):
#> rowLinks: a LinkDataFrame (19216 rows)
#> rowTree: 1 phylo tree(s) (19216 leaves)
#> colLinks: NULL
#> colTree: NULL
merged <- agglomerateByVariable(
GlobalPatterns, by = "cols", colData(GlobalPatterns)$SampleType)
merged
#> class: TreeSummarizedExperiment
#> dim: 19216 9
#> metadata(0):
#> assays(1): counts
#> rownames(19216): 549322 522457 ... 200359 271582
#> rowData names(7): Kingdom Phylum ... Genus Species
#> colnames(9): Feces Freshwater ... Soil Tongue
#> colData names(7): X.SampleID Primer ... SampleType Description
#> reducedDimNames(0):
#> mainExpName: NULL
#> altExpNames(0):
#> rowLinks: a LinkDataFrame (19216 rows)
#> rowTree: 1 phylo tree(s) (19216 leaves)
#> colLinks: NULL
#> colTree: NULL
## Agglomerate by multiple modules
# Generate 30 random modules
N_module <- 30L
modules <- sample(
c(TRUE, FALSE),
size = nrow(tse) * N_module,
prob = c(0.2, 0.8),
replace = TRUE
)
# Convert modules to matrix
modules <- matrix(modules, nrow = nrow(tse))
# Add module names as colnames
colnames(modules) <- paste0("module_", seq_len(ncol(modules)))
# Add modules to rowData
rowData(tse) <- cbind(rowData(tse), modules)
# Extract module columns
module_columns <- grep("module_", colnames(rowData(tse)), value = TRUE)
# Agglomerate based on modules
tse_module <- agglomerateByModule(tse, by = 1, group = module_columns)
#> Warning: 'pa' includes binary values.
#> Agglomeration of it might lead to meaningless values.
#> Check the assay, and consider doing transformation againmanually with agglomerated data.
# Optionally, store results into altExp slot
altExp(tse, "modules") <- tse_module
data(GlobalPatterns)
# print the available taxonomic ranks
taxonomyRanks(GlobalPatterns)
#> [1] "Kingdom" "Phylum" "Class" "Order" "Family" "Genus" "Species"
# agglomerateByRanks
#
tse <- agglomerateByRanks(GlobalPatterns)
#> Duplicated labels were made unique.
#> Duplicated labels were made unique.
altExps(tse)
#> List of length 7
#> names(7): Kingdom Phylum Class Order Family Genus Species
altExp(tse,"Kingdom")
#> class: TreeSummarizedExperiment
#> dim: 2 26
#> metadata(1): agglomerated_by_rank
#> assays(1): counts
#> rownames(2): Archaea Bacteria
#> rowData names(7): Kingdom Phylum ... Genus Species
#> colnames(26): CL3 CC1 ... Even2 Even3
#> colData names(7): X.SampleID Primer ... SampleType Description
#> reducedDimNames(0):
#> mainExpName: NULL
#> altExpNames(0):
#> rowLinks: a LinkDataFrame (2 rows)
#> rowTree: 1 phylo tree(s) (2 leaves)
#> colLinks: NULL
#> colTree: NULL
altExp(tse,"Species")
#> class: TreeSummarizedExperiment
#> dim: 944 26
#> metadata(1): agglomerated_by_rank
#> assays(1): counts
#> rownames(944): Abiotrophiadefectiva Achromatiumoxaliferum ...
#> proteobacteriumsymbiontofOsedaxsp.MB4 symbiontofNoeetapupillata
#> rowData names(7): Kingdom Phylum ... Genus Species
#> colnames(26): CL3 CC1 ... Even2 Even3
#> colData names(7): X.SampleID Primer ... SampleType Description
#> reducedDimNames(0):
#> mainExpName: NULL
#> altExpNames(0):
#> rowLinks: a LinkDataFrame (944 rows)
#> rowTree: 1 phylo tree(s) (944 leaves)
#> colLinks: NULL
#> colTree: NULL
# unsplitByRanks
tse <- unsplitByRanks(tse)
tse
#> class: TreeSummarizedExperiment
#> dim: 2692 26
#> metadata(0):
#> assays(1): counts
#> rownames(2692): Kingdom:Archaea Kingdom:Bacteria ...
#> Species:proteobacteriumsymbiontofOsedaxsp.MB4
#> Species:symbiontofNoeetapupillata
#> rowData names(8): Kingdom Phylum ... Species taxonomicLevel
#> colnames(26): CL3 CC1 ... Even2 Even3
#> colData names(7): X.SampleID Primer ... SampleType Description
#> reducedDimNames(0):
#> mainExpName: NULL
#> altExpNames(0):
#> rowLinks: NULL
#> rowTree: NULL
#> colLinks: NULL
#> colTree: NULL