plotForest() creates a feature- or sample-wise forest plot, showing
estimated results from a statistical test with their confidence intervals.
Additionally, the plot can be enriched with the tree structure and labelled
with Confidence Intervals (CIs), p-values and other side information.
plotForest(x, ...)
# S4 method for class 'TreeSummarizedExperiment'
plotForest(
x,
by = 1L,
effect.var = "effect",
ci.lower.var = "lower",
ci.upper.var = "upper",
err.var = NULL,
pval.var = "pval",
id.var = "rownames",
label.by = NULL,
order.by = NULL,
facet.by = NULL,
color.by = colour.by,
colour.by = NULL,
conf.level = 0.95,
tree.name = "phylo",
show.tree = TRUE,
...
)
# S4 method for class 'SummarizedExperiment'
plotForest(
x,
by = 1L,
effect.var = "effect",
ci.lower.var = "lower",
ci.upper.var = "upper",
err.var = NULL,
pval.var = "pval",
id.var = "rownames",
label.by = NULL,
order.by = NULL,
facet.by = NULL,
color.by = colour.by,
colour.by = NULL,
conf.level = 0.95
)
# S4 method for class 'data.frame'
plotForest(
x,
effect.var = "effect",
ci.lower.var = "lower",
ci.upper.var = "upper",
err.var = NULL,
pval.var = "pval",
id.var = "rownames",
label.by = NULL,
order.by = NULL,
facet.by = NULL,
color.by = colour.by,
colour.by = NULL,
conf.level = 0.95
)Arguments
- x
a
SummarizedExperimentobject, or adata.frameobject containing statistical estimates.- ...
additional parameters passed to
plotRowTree.- by
Character scalar. Determines whether features or samples data is used for the plot. (Default:"rows")- effect.var
Character scalar. Specifies the variable of x which corresponds to the effects or estimated results. (Default:"effect")- ci.lower.var
Character scalar. Specifies the variable of x which corresponds to the lower CI boundaries. (Default:"lower")- ci.upper.var
Character scalar. Specifies the variable of x which corresponds to the upper CI boundaries. (Default:"upper")- err.var
Character scalar. Specifies the variable of x which corresponds to the standard errors associated witheffect.var. When defined, it overwritesci.lower.varandci.upper.var. (Default:"pval")- pval.var
Character scalar. Specifies the variable of x which corresponds to the p-values associated witheffect.var. (Default:"pval")- id.var
Character scalar. Specifies the variable of x which corresponds to the observation identifiers. When"rownames"), the object rownames are used. (Default:"rownames")- label.by
Character vector. Specifies the variables of x or row/colData(x) by which the plot should be labelled."CI"and"P-Value"are special entries which require eithereffect.var,ci.lower.varandci.upper.varorpval.varto be specified, respectively. (Default:NULL)- order.by
Character scalar. Specifies the variable of x by which observations are ordered. IfNULL, the observations are ordered by the tree structure if available. (Default:NULL)- facet.by
Character scalar. Specifies the variable of x by which observations are divided into horizontal facets. (Default:NULL)- color.by
Character scalar. Alias tocolour.by.- colour.by
Character scalar. Specifies the variable of x by which observations are coloured. (Default:NULL)- conf.level
Numeric scalar. Specifies the confidence level of the interval when inferred fromerr.var. It is ignored whenci.lower.varandci.upper.varare defined. (Default:0.95)- tree.name
Character scalar. Specifies a row/colTree from x. (Default:"phylo")- show.tree
Logical scalar. Should the tree structure of the data be shown next to the forest plot?
Value
a ggplot object.
Examples
library(mia)
library(maaslin3)
# Import dataset
data("Tengeler2020", package = "mia")
tse <- Tengeler2020
# Agglomerate by genus and subset by prevalence
tse <- subsetByPrevalent(tse, rank = "Genus", prevalence = 10/100)
#> Duplicated labels were made unique.
# Transform count assay to relative abundances
tse <- transformAssay(tse, assay.type = "counts", method = "relabundance")
# Run maaslin3
maaslin3_out <- maaslin3(
input_data = tse,
output = "maaslin_results",
formula = "~ patient_status",
)
#> 2026-06-01 07:53:38.96 INFO::Writing function arguments to log file
#> 2026-06-01 07:53:38.99 INFO::Verifying options selected are valid
#> 2026-06-01 07:53:38.99 INFO::Determining format of input files
#> 2026-06-01 07:53:38.99 INFO::Input format is data samples as rows and metadata samples as rows
#> 2026-06-01 07:53:38.99 INFO::Running selected normalization method: TSS
#> 2026-06-01 07:53:39.00 INFO::Creating output feature tables folder
#> 2026-06-01 07:53:39.00 INFO::Writing normalized data to file maaslin_results/features/data_norm.tsv
#> 2026-06-01 07:53:39.00 INFO::Filter data based on min abundance, min prevalence, and max prevalence
#> 2026-06-01 07:53:39.00 INFO::Total samples in data: 27
#> 2026-06-01 07:53:39.00 INFO::Min samples required with min abundance for a feature not to be filtered: 0.000000
#> 2026-06-01 07:53:39.00 INFO::Max samples allowed with min abundance for a feature not to be filtered: 27.270000
#> 2026-06-01 07:53:39.00 INFO::Total filtered features: 0
#> 2026-06-01 07:53:39.00 INFO::Filtered feature names from abundance, min prevalence, and max prevalence filtering:
#> 2026-06-01 07:53:39.01 INFO::Total features filtered by non-zero variance filtering: 0
#> 2026-06-01 07:53:39.01 INFO::Filtered feature names from variance filtering:
#> 2026-06-01 07:53:39.01 INFO::Writing filtered data to file maaslin_results/features/filtered_data.tsv
#> 2026-06-01 07:53:39.01 INFO::Running selected transform method: LOG
#> 2026-06-01 07:53:39.01 INFO::Writing normalized, filtered, transformed data to file maaslin_results/features/data_transformed.tsv
#> 2026-06-01 07:53:39.01 INFO::Applying z-score to standardize continuous metadata
#> 2026-06-01 07:53:39.03 INFO::Running the linear model component
#> 2026-06-01 07:53:39.04 INFO::Fitting model to feature number 1, Akkermansia
#> 2026-06-01 07:53:39.04 INFO::Fitting model to feature number 2, Alistipes
#> 2026-06-01 07:53:39.05 INFO::Fitting model to feature number 3, Anaerostipes
#> 2026-06-01 07:53:39.05 INFO::Fitting model to feature number 4, Anaerotruncus
#> 2026-06-01 07:53:39.05 INFO::Fitting model to feature number 5, Bacteroides
#> 2026-06-01 07:53:39.06 INFO::Fitting model to feature number 6, Barnesiella
#> 2026-06-01 07:53:39.06 INFO::Fitting model to feature number 7, Blautia
#> 2026-06-01 07:53:39.06 INFO::Fitting model to feature number 8, Butyricicoccus
#> 2026-06-01 07:53:39.07 WARNING::Fitting problem for feature 8 returning NA
#> 2026-06-01 07:53:39.07 INFO::Fitting model to feature number 9, Catabacter
#> 2026-06-01 07:53:39.07 INFO::Fitting model to feature number 10, Clostridium_sensu_stricto_1
#> 2026-06-01 07:53:39.07 INFO::Fitting model to feature number 11, Coprobacter
#> 2026-06-01 07:53:39.07 INFO::Fitting model to feature number 12, Coprococcus_2
#> 2026-06-01 07:53:39.08 INFO::Fitting model to feature number 13, Dielma
#> 2026-06-01 07:53:39.08 INFO::Fitting model to feature number 14, Eisenbergiella
#> 2026-06-01 07:53:39.08 INFO::Fitting model to feature number 15, Epulopiscium
#> 2026-06-01 07:53:39.09 INFO::Fitting model to feature number 16, Erysipelatoclostridium
#> 2026-06-01 07:53:39.09 INFO::Fitting model to feature number 17, Escherichia-Shigella
#> 2026-06-01 07:53:39.09 INFO::Fitting model to feature number 18, Eubacterium
#> 2026-06-01 07:53:39.10 INFO::Fitting model to feature number 19, Faecalibacterium
#> 2026-06-01 07:53:39.10 INFO::Fitting model to feature number 20, Flavonifractor
#> 2026-06-01 07:53:39.10 INFO::Fitting model to feature number 21, Fusicatenibacter
#> 2026-06-01 07:53:39.11 INFO::Fitting model to feature number 22, Halomonas
#> 2026-06-01 07:53:39.11 INFO::Fitting model to feature number 23, Holdemania
#> 2026-06-01 07:53:39.11 INFO::Fitting model to feature number 24, Hungatella
#> 2026-06-01 07:53:39.12 INFO::Fitting model to feature number 25, Lachnoclostridium
#> 2026-06-01 07:53:39.12 INFO::Fitting model to feature number 26, Lachnospiraceae_ND3007_group
#> 2026-06-01 07:53:39.12 INFO::Fitting model to feature number 27, Odoribacter
#> 2026-06-01 07:53:39.13 INFO::Fitting model to feature number 28, Parabacteroides
#> 2026-06-01 07:53:39.13 INFO::Fitting model to feature number 29, Paraprevotella
#> 2026-06-01 07:53:39.13 INFO::Fitting model to feature number 30, Roseburia
#> 2026-06-01 07:53:39.14 INFO::Fitting model to feature number 31, Ruminiclostridium_5
#> 2026-06-01 07:53:39.14 INFO::Fitting model to feature number 32, Ruminiclostridium_9
#> 2026-06-01 07:53:39.14 INFO::Fitting model to feature number 33, Ruminococcaceae_UCG-004
#> 2026-06-01 07:53:39.15 INFO::Fitting model to feature number 34, Ruminococcaceae_UCG-013
#> 2026-06-01 07:53:39.15 INFO::Fitting model to feature number 35, Ruminococcaceae_UCG-014
#> 2026-06-01 07:53:39.15 WARNING::Fitting problem for feature 35 returning NA
#> 2026-06-01 07:53:39.15 INFO::Fitting model to feature number 36, Ruminococcus_1
#> 2026-06-01 07:53:39.15 INFO::Fitting model to feature number 37, Ruminococcus_2
#> 2026-06-01 07:53:39.16 INFO::Fitting model to feature number 38, Subdoligranulum
#> 2026-06-01 07:53:39.16 INFO::Fitting model to feature number 39, [Clostridium]_innocuum_group
#> 2026-06-01 07:53:39.16 INFO::Fitting model to feature number 40, [Eubacterium]_coprostanoligenes_group
#> 2026-06-01 07:53:39.17 INFO::Fitting model to feature number 41, [Eubacterium]_fissicatena_group
#> 2026-06-01 07:53:39.17 INFO::Fitting model to feature number 42, [Eubacterium]_rectale_group
#> 2026-06-01 07:53:39.17 INFO::Fitting model to feature number 43, [Eubacterium]_xylanophilum_group
#> 2026-06-01 07:53:39.18 INFO::Fitting model to feature number 44, [Ruminococcus]_gauvreauii_group
#> 2026-06-01 07:53:39.18 INFO::Fitting model to feature number 45, [Ruminococcus]_gnavus_group
#> 2026-06-01 07:53:39.19 INFO::Fitting model to feature number 46, uncultured
#> 2026-06-01 07:53:39.19 INFO::Fitting model to feature number 47, uncultured_1
#> 2026-06-01 07:53:39.19 INFO::Fitting model to feature number 48, uncultured_2
#> 2026-06-01 07:53:39.19 INFO::Fitting model to feature number 49, uncultured_bacterium
#> 2026-06-01 07:53:39.23 INFO::Performing tests against medians
#> 2026-06-01 07:53:39.66 INFO::Counting total values for each feature
#> 2026-06-01 07:53:39.66 INFO::Running the logistic model component
#> 2026-06-01 07:53:39.67 INFO::Fitting model to feature number 1, Akkermansia
#> 2026-06-01 07:53:39.67 INFO::Fitting model to feature number 2, Alistipes
#> 2026-06-01 07:53:39.67 INFO::Fitting model to feature number 3, Anaerostipes
#> 2026-06-01 07:53:39.67 INFO::Fitting model to feature number 4, Anaerotruncus
#> 2026-06-01 07:53:39.68 INFO::Fitting model to feature number 5, Bacteroides
#> 2026-06-01 07:53:39.68 INFO::Fitting model to feature number 6, Barnesiella
#> 2026-06-01 07:53:39.69 INFO::Fitting model to feature number 7, Blautia
#> 2026-06-01 07:53:39.69 INFO::Fitting model to feature number 8, Butyricicoccus
#> 2026-06-01 07:53:39.70 INFO::Fitting model to feature number 9, Catabacter
#> 2026-06-01 07:53:39.70 INFO::Fitting model to feature number 10, Clostridium_sensu_stricto_1
#> 2026-06-01 07:53:39.70 INFO::Fitting model to feature number 11, Coprobacter
#> 2026-06-01 07:53:39.71 INFO::Fitting model to feature number 12, Coprococcus_2
#> 2026-06-01 07:53:39.71 INFO::Fitting model to feature number 13, Dielma
#> 2026-06-01 07:53:39.72 INFO::Fitting model to feature number 14, Eisenbergiella
#> 2026-06-01 07:53:39.72 INFO::Fitting model to feature number 15, Epulopiscium
#> 2026-06-01 07:53:39.73 INFO::Fitting model to feature number 16, Erysipelatoclostridium
#> 2026-06-01 07:53:39.73 INFO::Fitting model to feature number 17, Escherichia-Shigella
#> 2026-06-01 07:53:39.73 INFO::Fitting model to feature number 18, Eubacterium
#> 2026-06-01 07:53:39.74 INFO::Fitting model to feature number 19, Faecalibacterium
#> 2026-06-01 07:53:39.74 INFO::Fitting model to feature number 20, Flavonifractor
#> 2026-06-01 07:53:39.75 INFO::Fitting model to feature number 21, Fusicatenibacter
#> 2026-06-01 07:53:39.75 INFO::Fitting model to feature number 22, Halomonas
#> 2026-06-01 07:53:39.76 INFO::Fitting model to feature number 23, Holdemania
#> 2026-06-01 07:53:39.76 INFO::Fitting model to feature number 24, Hungatella
#> 2026-06-01 07:53:39.76 INFO::Fitting model to feature number 25, Lachnoclostridium
#> 2026-06-01 07:53:39.77 INFO::Fitting model to feature number 26, Lachnospiraceae_ND3007_group
#> 2026-06-01 07:53:39.77 INFO::Fitting model to feature number 27, Odoribacter
#> 2026-06-01 07:53:39.77 INFO::Fitting model to feature number 28, Parabacteroides
#> 2026-06-01 07:53:39.78 INFO::Fitting model to feature number 29, Paraprevotella
#> 2026-06-01 07:53:39.78 INFO::Fitting model to feature number 30, Roseburia
#> 2026-06-01 07:53:39.79 INFO::Fitting model to feature number 31, Ruminiclostridium_5
#> 2026-06-01 07:53:39.79 INFO::Fitting model to feature number 32, Ruminiclostridium_9
#> 2026-06-01 07:53:39.80 INFO::Fitting model to feature number 33, Ruminococcaceae_UCG-004
#> 2026-06-01 07:53:39.80 INFO::Fitting model to feature number 34, Ruminococcaceae_UCG-013
#> 2026-06-01 07:53:39.80 INFO::Fitting model to feature number 35, Ruminococcaceae_UCG-014
#> 2026-06-01 07:53:39.80 INFO::Fitting model to feature number 36, Ruminococcus_1
#> 2026-06-01 07:53:39.81 INFO::Fitting model to feature number 37, Ruminococcus_2
#> 2026-06-01 07:53:39.81 INFO::Fitting model to feature number 38, Subdoligranulum
#> 2026-06-01 07:53:39.82 INFO::Fitting model to feature number 39, [Clostridium]_innocuum_group
#> 2026-06-01 07:53:39.82 INFO::Fitting model to feature number 40, [Eubacterium]_coprostanoligenes_group
#> 2026-06-01 07:53:39.83 INFO::Fitting model to feature number 41, [Eubacterium]_fissicatena_group
#> 2026-06-01 07:53:39.83 INFO::Fitting model to feature number 42, [Eubacterium]_rectale_group
#> 2026-06-01 07:53:39.83 INFO::Fitting model to feature number 43, [Eubacterium]_xylanophilum_group
#> 2026-06-01 07:53:39.84 INFO::Fitting model to feature number 44, [Ruminococcus]_gauvreauii_group
#> 2026-06-01 07:53:39.84 INFO::Fitting model to feature number 45, [Ruminococcus]_gnavus_group
#> 2026-06-01 07:53:39.85 INFO::Fitting model to feature number 46, uncultured
#> 2026-06-01 07:53:39.85 INFO::Fitting model to feature number 47, uncultured_1
#> 2026-06-01 07:53:39.86 INFO::Fitting model to feature number 48, uncultured_2
#> 2026-06-01 07:53:39.86 INFO::Fitting model to feature number 49, uncultured_bacterium
#> 2026-06-01 07:53:39.87 INFO::Counting total values for each feature
#> 2026-06-01 07:53:39.87 INFO::Re-running abundances for warn_prevalence
#> 2026-06-01 07:53:39.87 INFO::Running selected normalization method: TSS
#> 2026-06-01 07:53:39.87 INFO::Running selected transform method: LOG
#> 2026-06-01 07:53:39.88 INFO::Fitting model to feature number 1, Akkermansia
#> 2026-06-01 07:53:39.88 INFO::Fitting model to feature number 2, Alistipes
#> 2026-06-01 07:53:39.88 INFO::Fitting model to feature number 3, Anaerostipes
#> 2026-06-01 07:53:39.89 INFO::Fitting model to feature number 4, Anaerotruncus
#> 2026-06-01 07:53:39.89 INFO::Fitting model to feature number 5, Bacteroides
#> 2026-06-01 07:53:39.89 INFO::Fitting model to feature number 6, Barnesiella
#> 2026-06-01 07:53:39.90 INFO::Fitting model to feature number 7, Blautia
#> 2026-06-01 07:53:39.90 INFO::Fitting model to feature number 8, Butyricicoccus
#> 2026-06-01 07:53:39.90 WARNING::Fitting problem for feature 8 returning NA
#> 2026-06-01 07:53:39.90 INFO::Fitting model to feature number 9, Catabacter
#> 2026-06-01 07:53:39.91 INFO::Fitting model to feature number 10, Clostridium_sensu_stricto_1
#> 2026-06-01 07:53:39.91 INFO::Fitting model to feature number 11, Coprobacter
#> 2026-06-01 07:53:39.91 INFO::Fitting model to feature number 12, Coprococcus_2
#> 2026-06-01 07:53:39.91 INFO::Fitting model to feature number 13, Dielma
#> 2026-06-01 07:53:39.92 INFO::Fitting model to feature number 14, Eisenbergiella
#> 2026-06-01 07:53:39.92 INFO::Fitting model to feature number 15, Epulopiscium
#> 2026-06-01 07:53:39.92 INFO::Fitting model to feature number 16, Erysipelatoclostridium
#> 2026-06-01 07:53:39.93 INFO::Fitting model to feature number 17, Escherichia-Shigella
#> 2026-06-01 07:53:39.93 INFO::Fitting model to feature number 18, Eubacterium
#> 2026-06-01 07:53:39.93 INFO::Fitting model to feature number 19, Faecalibacterium
#> 2026-06-01 07:53:39.93 INFO::Fitting model to feature number 20, Flavonifractor
#> 2026-06-01 07:53:39.94 INFO::Fitting model to feature number 21, Fusicatenibacter
#> 2026-06-01 07:53:39.94 INFO::Fitting model to feature number 22, Halomonas
#> 2026-06-01 07:53:39.94 INFO::Fitting model to feature number 23, Holdemania
#> 2026-06-01 07:53:39.95 INFO::Fitting model to feature number 24, Hungatella
#> 2026-06-01 07:53:39.95 INFO::Fitting model to feature number 25, Lachnoclostridium
#> 2026-06-01 07:53:39.95 INFO::Fitting model to feature number 26, Lachnospiraceae_ND3007_group
#> 2026-06-01 07:53:39.96 INFO::Fitting model to feature number 27, Odoribacter
#> 2026-06-01 07:53:39.96 INFO::Fitting model to feature number 28, Parabacteroides
#> 2026-06-01 07:53:39.96 INFO::Fitting model to feature number 29, Paraprevotella
#> 2026-06-01 07:53:39.97 INFO::Fitting model to feature number 30, Roseburia
#> 2026-06-01 07:53:39.97 INFO::Fitting model to feature number 31, Ruminiclostridium_5
#> 2026-06-01 07:53:39.97 INFO::Fitting model to feature number 32, Ruminiclostridium_9
#> 2026-06-01 07:53:39.97 INFO::Fitting model to feature number 33, Ruminococcaceae_UCG-004
#> 2026-06-01 07:53:39.98 INFO::Fitting model to feature number 34, Ruminococcaceae_UCG-013
#> 2026-06-01 07:53:39.98 INFO::Fitting model to feature number 35, Ruminococcaceae_UCG-014
#> 2026-06-01 07:53:39.98 WARNING::Fitting problem for feature 35 returning NA
#> 2026-06-01 07:53:39.98 INFO::Fitting model to feature number 36, Ruminococcus_1
#> 2026-06-01 07:53:39.99 INFO::Fitting model to feature number 37, Ruminococcus_2
#> 2026-06-01 07:53:39.99 INFO::Fitting model to feature number 38, Subdoligranulum
#> 2026-06-01 07:53:39.99 INFO::Fitting model to feature number 39, [Clostridium]_innocuum_group
#> 2026-06-01 07:53:40.00 INFO::Fitting model to feature number 40, [Eubacterium]_coprostanoligenes_group
#> 2026-06-01 07:53:40.00 INFO::Fitting model to feature number 41, [Eubacterium]_fissicatena_group
#> 2026-06-01 07:53:40.00 INFO::Fitting model to feature number 42, [Eubacterium]_rectale_group
#> 2026-06-01 07:53:40.00 INFO::Fitting model to feature number 43, [Eubacterium]_xylanophilum_group
#> 2026-06-01 07:53:40.01 INFO::Fitting model to feature number 44, [Ruminococcus]_gauvreauii_group
#> 2026-06-01 07:53:40.01 INFO::Fitting model to feature number 45, [Ruminococcus]_gnavus_group
#> 2026-06-01 07:53:40.01 INFO::Fitting model to feature number 46, uncultured
#> 2026-06-01 07:53:40.02 INFO::Fitting model to feature number 47, uncultured_1
#> 2026-06-01 07:53:40.02 INFO::Fitting model to feature number 48, uncultured_2
#> 2026-06-01 07:53:40.02 INFO::Fitting model to feature number 49, uncultured_bacterium
#> 2026-06-01 07:53:40.03 INFO::Creating fits folder
#> 2026-06-01 07:53:40.03 INFO::Writing residuals to file maaslin_results/fits/residuals_linear.rds
#> 2026-06-01 07:53:40.03 INFO::Writing fitted values to file maaslin_results/fits/fitted_linear.rds
#> 2026-06-01 07:53:40.04 INFO::Writing residuals to file maaslin_results/fits/residuals_logistic.rds
#> 2026-06-01 07:53:40.04 INFO::Writing fitted values to file maaslin_results/fits/fitted_logistic.rds
#> 2026-06-01 07:53:40.04 INFO::Writing all the results to file (ordered
#> by increasing individual q-values): maaslin_results/all_results.tsv
#> 2026-06-01 07:53:40.04 INFO::Writing the significant results without errors (those which have joint q-values less than or equal to the threshold of 0.100000 ) to file (ordered by increasing individual q-values): maaslin_results/significant_results.tsv
#> 2026-06-01 07:53:40.04 INFO::Creating output figures folder
#> 2026-06-01 07:53:40.04 INFO::Writing summary plot of significant
#> results to file: maaslin_results/figures/summary_plot.pdf
#> 2026-06-01 07:53:41.23 INFO::Writing association plots (one for each significant association) to output folder: maaslin_results/figures
#> 2026-06-01 07:53:41.24 INFO::Plotting associations from most to least significant, grouped by metadata
#> 2026-06-01 07:53:41.24 INFO::Creating box plot for categorical data (linear),
#> patient_status vs Coprobacter
#> 2026-06-01 07:53:41.61 INFO::Creating box plot for categorical data (linear),
#> patient_status vs Faecalibacterium
#> 2026-06-01 07:53:41.93 INFO::Creating tile plot for categorical data (logistic), patient_status vs Erysipelatoclostridium
# Retrieve abundance results
maaslin3_abund <- maaslin3_out$fit_data_abundance$results
maaslin3_abund <- maaslin3_abund[!is.na(maaslin3_abund$coef), ]
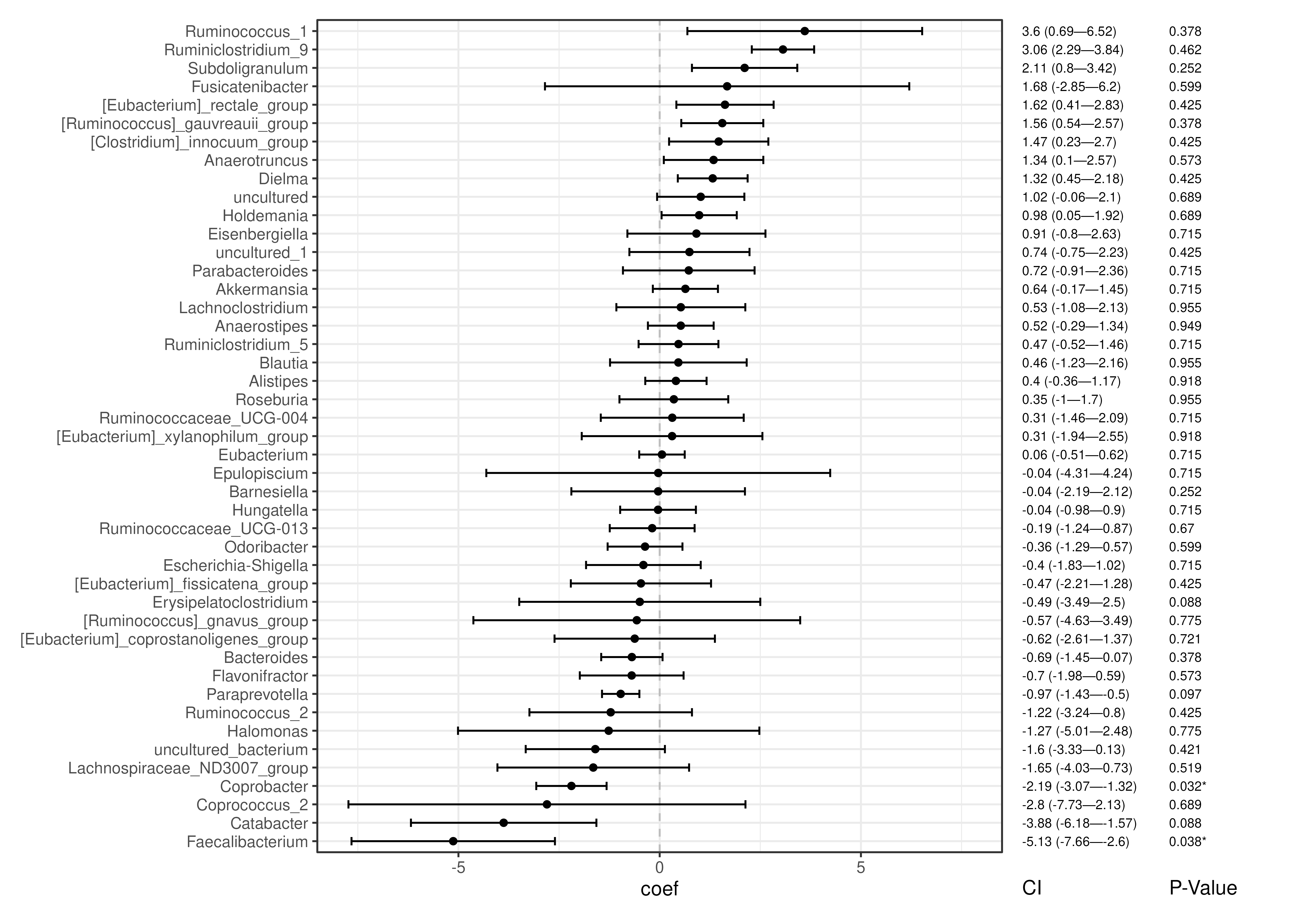
# Visualize abundance results
plotForest(
maaslin3_abund,
effect.var = "coef",
err.var = "stderr",
pval.var = "qval_joint",
id.var = "feature",
label.by = c("CI", "P-Value"),
order.by = "coef"
)
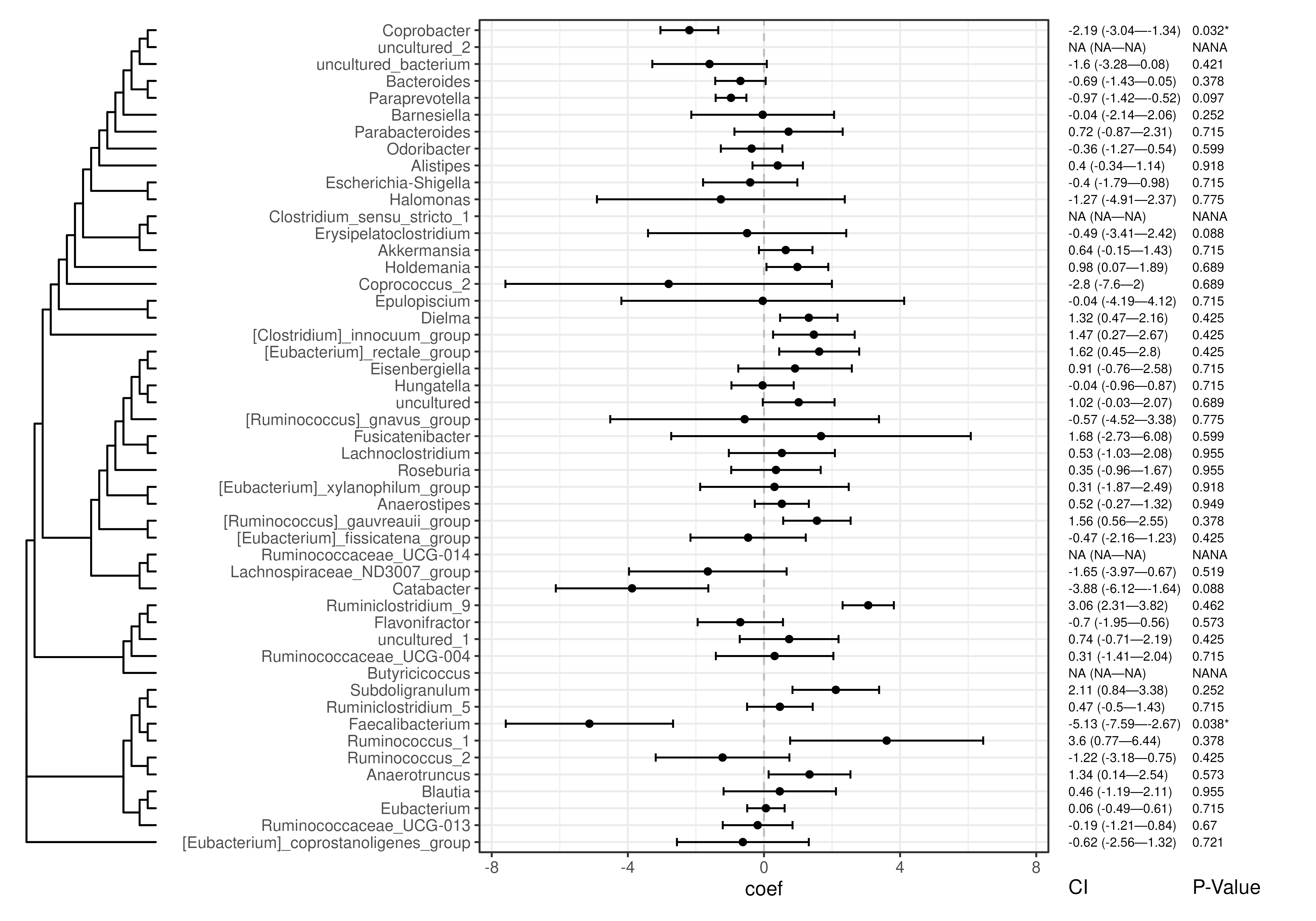
 # Add abundance results to TreeSE rowData
rownames(maaslin3_abund) <- maaslin3_abund$feature
tax_order <- match(rownames(tse), rownames(maaslin3_abund))
rowData(tse) <- cbind(rowData(tse), maaslin3_abund[tax_order, ])
# Visualise abundance results with tree structure
plotForest(
tse,
effect.var = "coef",
err.var = "stderr",
pval.var = "qval_joint",
label.by = c("CI", "P-Value")
)
#> Warning: `aes_string()` was deprecated in ggplot2 3.0.0.
#> ℹ Please use tidy evaluation idioms with `aes()`.
#> ℹ See also `vignette("ggplot2-in-packages")` for more information.
#> ℹ The deprecated feature was likely used in the miaViz package.
#> Please report the issue at <https://github.com/microbiome/miaViz/issues>.
#> Warning: Removed 4 rows containing missing values or values outside the scale range
#> (`geom_point()`).
# Add abundance results to TreeSE rowData
rownames(maaslin3_abund) <- maaslin3_abund$feature
tax_order <- match(rownames(tse), rownames(maaslin3_abund))
rowData(tse) <- cbind(rowData(tse), maaslin3_abund[tax_order, ])
# Visualise abundance results with tree structure
plotForest(
tse,
effect.var = "coef",
err.var = "stderr",
pval.var = "qval_joint",
label.by = c("CI", "P-Value")
)
#> Warning: `aes_string()` was deprecated in ggplot2 3.0.0.
#> ℹ Please use tidy evaluation idioms with `aes()`.
#> ℹ See also `vignette("ggplot2-in-packages")` for more information.
#> ℹ The deprecated feature was likely used in the miaViz package.
#> Please report the issue at <https://github.com/microbiome/miaViz/issues>.
#> Warning: Removed 4 rows containing missing values or values outside the scale range
#> (`geom_point()`).
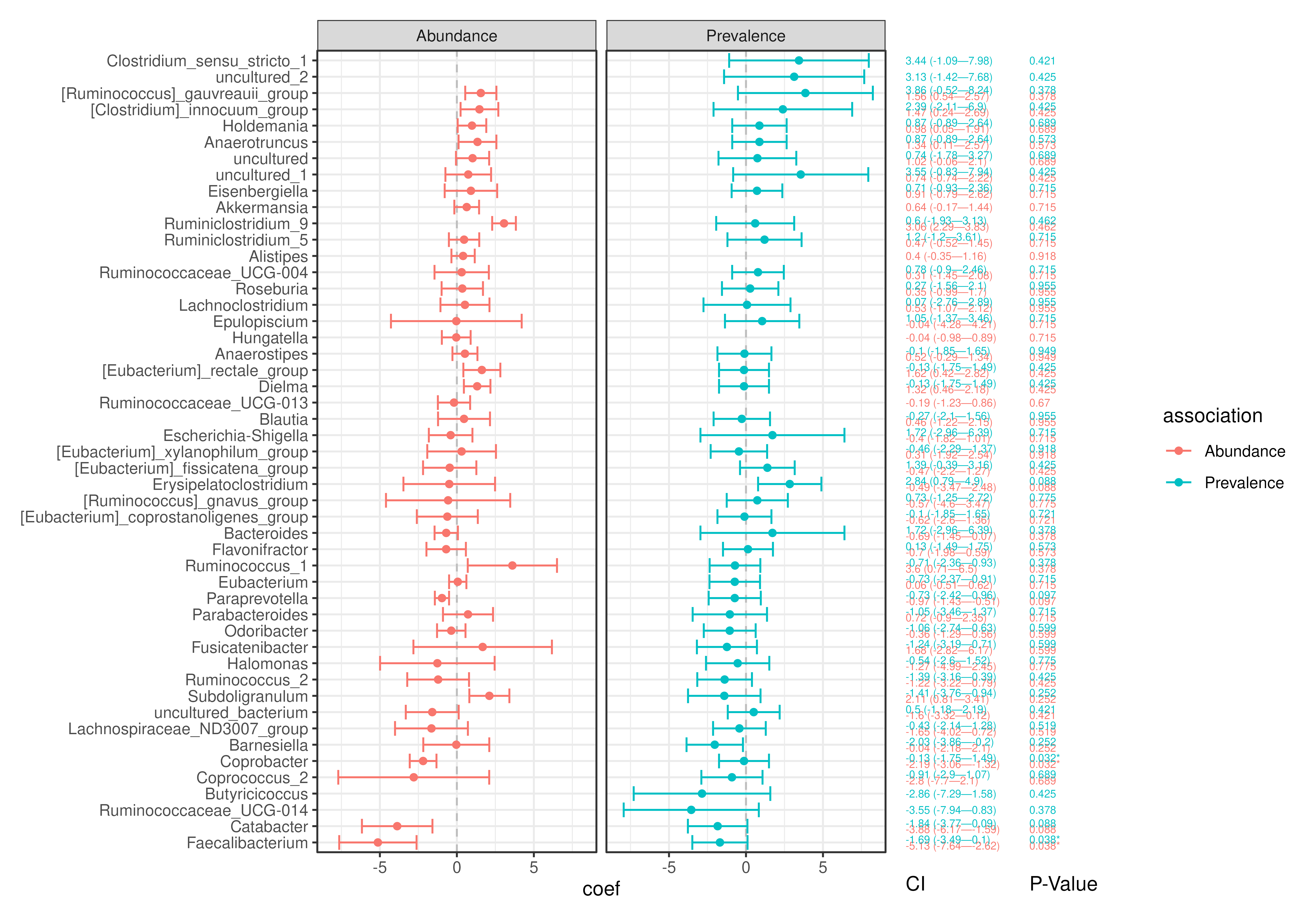
 # Retrieve prevalence results
maaslin3_prev <- maaslin3_out$fit_data_prevalence$results
maaslin3_prev <- maaslin3_prev[!is.na(maaslin3_prev$coef), ]
# Combine abundance and prevalence results
maaslin3_res <- rbind(maaslin3_abund, maaslin3_prev)
maaslin3_res$association <- c(
rep("Abundance", nrow(maaslin3_abund)),
rep("Prevalence", nrow(maaslin3_prev))
)
# Visualize combined results
plotForest(
maaslin3_res,
effect.var = "coef",
err.var = "stderr",
pval.var = "qval_joint",
id.var = "feature",
label.by = c("CI", "P-Value"),
order.by = "coef",
facet.by = "association",
colour.by = "association"
)
# Retrieve prevalence results
maaslin3_prev <- maaslin3_out$fit_data_prevalence$results
maaslin3_prev <- maaslin3_prev[!is.na(maaslin3_prev$coef), ]
# Combine abundance and prevalence results
maaslin3_res <- rbind(maaslin3_abund, maaslin3_prev)
maaslin3_res$association <- c(
rep("Abundance", nrow(maaslin3_abund)),
rep("Prevalence", nrow(maaslin3_prev))
)
# Visualize combined results
plotForest(
maaslin3_res,
effect.var = "coef",
err.var = "stderr",
pval.var = "qval_joint",
id.var = "feature",
label.by = c("CI", "P-Value"),
order.by = "coef",
facet.by = "association",
colour.by = "association"
)